DepleteX® Single Cell RNA Boost Kit
Boost usable single cell RNA seq data with DepleteX®

This product is a new and improved replacement for CRISPRclean Single Cell RNA Boost Kit (KIT1018).
DepleteX® removes abundant and uninformative fragments before sequencing, redistributing 50% sequencing clusters to informative reads that normally would be filtered out prior to secondary analysis. This kit leverages Cas9 depletion with an optimized guide set to remove ribosomal and mitochondrial mRNA, non-transcriptomic reads, and highly expressed non-variable genes. This new kit includes a separate tube for non-variable gene depletion content that can be included or excluded as needed.
DepleteX Single Cell RNA Boost Kit gives you the ability to cut through the noise and maximize usable data in your single cell RNA sequencing.
Highlights
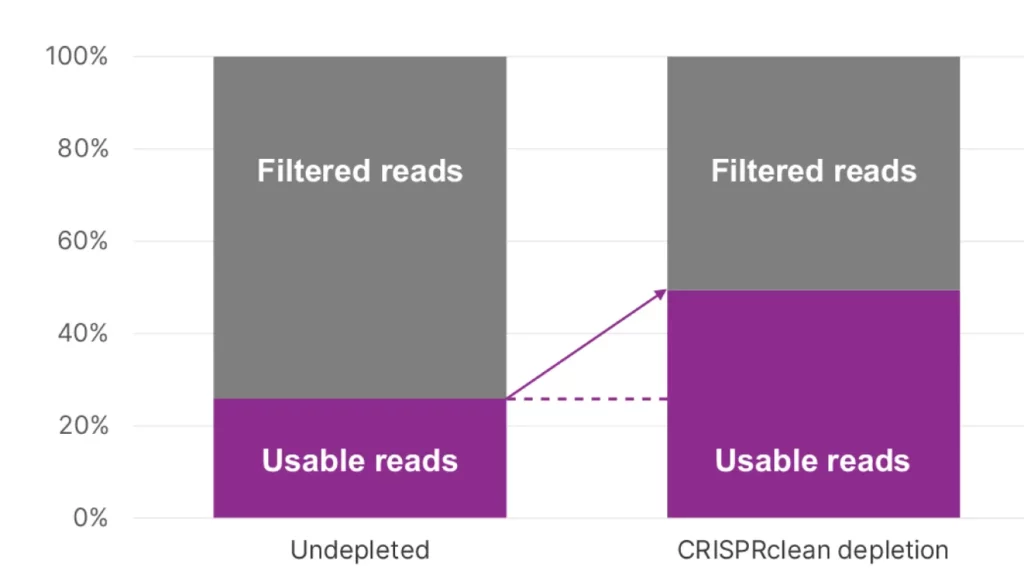
- ~50% boost in reads useful for secondary analysis
- Validated for both short and long read sequencing
- Depletes ribosomal, mitochondrial, non-transcriptomic reads, and non-variable genes (optional) from single cell libraries
- Simple 3 step protocol
| Specification | |
| Catalog | KIT1037 |
|---|---|
| Samples per kit | 24 |
| Assay time | 2 hours |
| Hands-on-time | 45 minutes |
| Input | Uses one of four Chromium cDNA aliquots per prep |
| Method | Single cell RNA-Seq |
| Designed to deplete | ribosomal, mitochondrial, non-transcriptomic reads, and non-variable genes (optional) |
| Jumpcode validated | 10x Genomics Chromium Next GEM Single Cell 3' Reagent Kits v3.1, Visium Spatial Gene Expression libraries, PacBio Iso-Seq SMRTbell Express Template Prep Kit 2.0, BD Rhapsody |
| Customer validated | 10x Genomics Chromium 5' and multiome, Parse Biosciences Evercode Whole Transcriptome |
Simple 3-step protocol integrated into 10x Genomics® Chromium™ Next GEM Single Cell 3’ workflow

100% increase in reads mapped to the transcriptome with DepleteX™ Single Cell RNA Boost Kit.
1.5x improvement in detection sensitivity with depletion
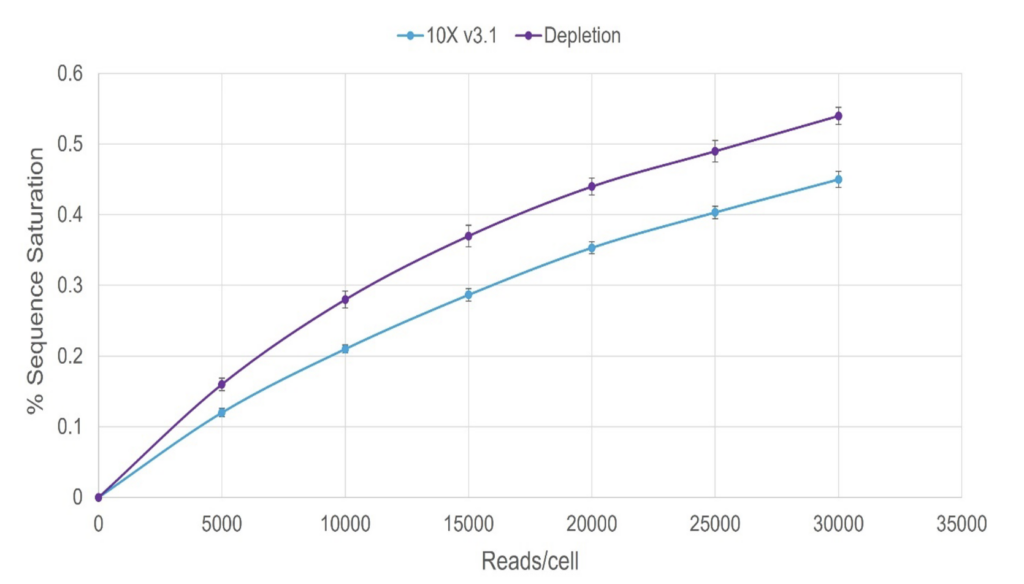
Boost in detection sensitivity of genes and UMIs per cell for depleted PBMC samples compared to undepleted sample.
Gain a deeper view of expression profiles
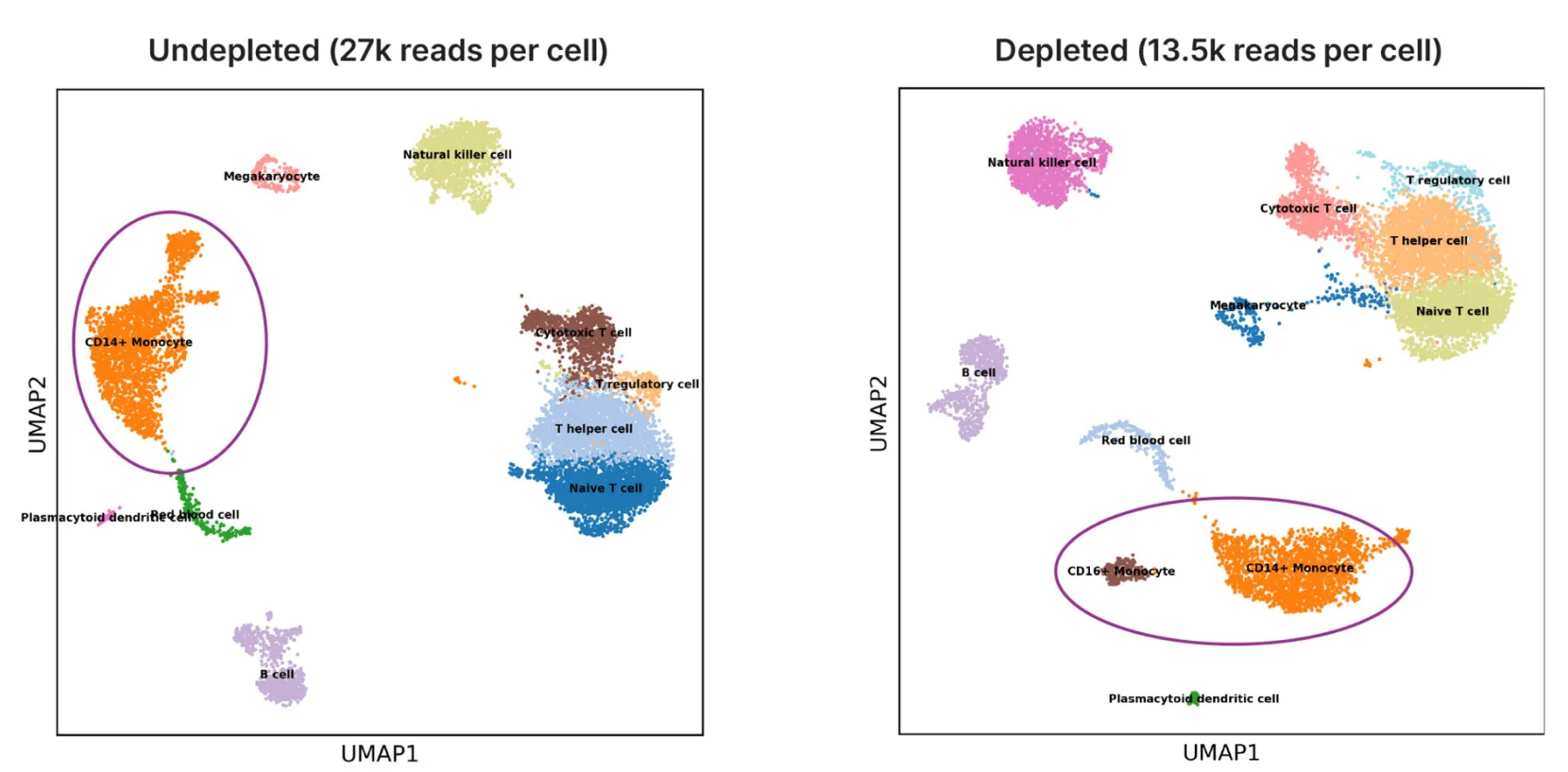
One additional cell type was identified in PBMC samples treated with DepleteX. UMAP plots of cell clusters with and without depletion using PBMC samples showed that depletion did not perturb cell type calls.

DepleteX® Single Cell RNA Boost Kit
24 samples
$6,000 Add to CartHow can we boost your single cell research?
Resources
Download helpful product documents.
Connect with us to learn more
For the US patent, the patent number is US 10,604,802 entitled Genome Fractioning. The patent publication is available here.
For the EP patent, the patent number is EP3102722 entitled Genome Fractioning. The patent publication is available here.